| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2022-03-07 02:48:59 UTC |
|---|
| HMDB ID | HMDB0000220 |
|---|
| Secondary Accession Numbers | - HMDB00220
- HMDB0060083
- HMDB60083
|
|---|
| Metabolite Identification |
|---|
| Common Name | Palmitic acid |
|---|
| Description | Palmitic acid, or hexadecanoic acid, is one of the most common saturated fatty acids found in animals, plants, and microorganisms. As its name indicates, it is a major component of the oil from the fruit of oil palms (palm oil). Excess carbohydrates in the body are converted to palmitic acid. Palmitic acid is the first fatty acid produced during fatty acid synthesis and is the precursor to longer fatty acids. As a consequence, palmitic acid is a major body component of animals. In humans, one analysis found it to make up 21–30% (molar) of human depot fat (PMID: 13756126 ), and it is a major, but highly variable, lipid component of human breast milk (PMID: 352132 ). Palmitic acid is used to produce soaps, cosmetics, and industrial mould release agents. These applications use sodium palmitate, which is commonly obtained by saponification of palm oil. To this end, palm oil, rendered from palm tree (species Elaeis guineensis), is treated with sodium hydroxide (in the form of caustic soda or lye), which causes hydrolysis of the ester groups, yielding glycerol and sodium palmitate. Aluminium salts of palmitic acid and naphthenic acid were combined during World War II to produce napalm. The word "napalm" is derived from the words naphthenic acid and palmitic acid (Wikipedia ). Palmitic acid is also used in the determination of water hardness and is a surfactant of Levovist, an intravenous ultrasonic contrast agent. |
|---|
| Structure | InChI=1S/C16H32O2/c1-2-3-4-5-6-7-8-9-10-11-12-13-14-15-16(17)18/h2-15H2,1H3,(H,17,18) |
|---|
| Synonyms | | Value | Source |
|---|
| 1-Hexyldecanoic acid | ChEBI | | 1-Pentadecanecarboxylic acid | ChEBI | | 16:00 | ChEBI | | C16 | ChEBI | | C16 Fatty acid | ChEBI | | C16:0 | ChEBI | | Cetylic acid | ChEBI | | CH3-[CH2]14-COOH | ChEBI | | FA 16:0 | ChEBI | | Hexadecanoate | ChEBI | | Hexadecoic acid | ChEBI | | Hexadecylic acid | ChEBI | | Hexaectylic acid | ChEBI | | N-Hexadecanoic acid | ChEBI | | N-Hexadecoic acid | ChEBI | | Palmitate | ChEBI | | Palmitinic acid | ChEBI | | Palmitinsaeure | ChEBI | | Pentadecanecarboxylic acid | ChEBI | | 1-Hexyldecanoate | Generator | | 1-Pentadecanecarboxylate | Generator | | Cetylate | Generator | | Hexadecanoic acid | Generator | | Hexadecoate | Generator | | Hexadecylate | Generator | | Hexaectylate | Generator | | N-Hexadecanoate | Generator | | N-Hexadecoate | Generator | | Palmitinate | Generator | | Pentadecanecarboxylate | Generator | | Acid, hexadecanoic | MeSH | | Acid, palmitic | MeSH | | Edenor C16 | HMDB | | Emersol 140 | HMDB | | Emersol 143 | HMDB | | Glycon P-45 | HMDB | | Hexadecanoate (N-C16:0) | HMDB | | Hexadecanoic acid palmitic acid | HMDB | | Hydrofol | HMDB | | Hydrofol acid 1690 | HMDB | | Hystrene 8016 | HMDB | | Hystrene 9016 | HMDB | | Industrene 4516 | HMDB | | Kortacid 1698 | HMDB | | Loxiol ep 278 | HMDB | | Lunac P 95 | HMDB | | Lunac P 95KC | HMDB | | Lunac P 98 | HMDB | | Palmitoate | HMDB | | Palmitoic acid | HMDB | | PAM | HMDB | | PLM | HMDB | | Prifac 2960 | HMDB | | Prifrac 2960 | HMDB | | Pristerene 4934 | HMDB | | Univol u332 | HMDB | | FA(16:0) | HMDB |
|
|---|
| Chemical Formula | C16H32O2 |
|---|
| Average Molecular Weight | 256.4241 |
|---|
| Monoisotopic Molecular Weight | 256.240230268 |
|---|
| IUPAC Name | hexadecanoic acid |
|---|
| Traditional Name | palmitic acid |
|---|
| CAS Registry Number | 57-10-3 |
|---|
| SMILES | CCCCCCCCCCCCCCCC(O)=O |
|---|
| InChI Identifier | InChI=1S/C16H32O2/c1-2-3-4-5-6-7-8-9-10-11-12-13-14-15-16(17)18/h2-15H2,1H3,(H,17,18) |
|---|
| InChI Key | IPCSVZSSVZVIGE-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as long-chain fatty acids. These are fatty acids with an aliphatic tail that contains between 13 and 21 carbon atoms. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Lipids and lipid-like molecules |
|---|
| Class | Fatty Acyls |
|---|
| Sub Class | Fatty acids and conjugates |
|---|
| Direct Parent | Long-chain fatty acids |
|---|
| Alternative Parents | |
|---|
| Substituents | - Long-chain fatty acid
- Straight chain fatty acid
- Monocarboxylic acid or derivatives
- Carboxylic acid
- Carboxylic acid derivative
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Carbonyl group
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Not Available | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | 61.8 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 4.0e-05 mg/mL | Not Available | | LogP | 7.17 | SANGSTER (1993) |
|
|---|
| Experimental Chromatographic Properties | Experimental Collision Cross Sections |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Retention Times Underivatized| Chromatographic Method | Retention Time | Reference |
|---|
| Measured using a Waters Acquity ultraperformance liquid chromatography (UPLC) ethylene-bridged hybrid (BEH) C18 column (100 mm × 2.1 mm; 1.7 μmparticle diameter). Predicted by Afia on May 17, 2022. Predicted by Afia on May 17, 2022. | 10.77 minutes | 32390414 | | Predicted by Siyang on May 30, 2022 | 22.9766 minutes | 33406817 | | Predicted by Siyang using ReTip algorithm on June 8, 2022 | 2.58 minutes | 32390414 | | AjsUoB = Accucore 150 Amide HILIC with 10mM Ammonium Formate, 0.1% Formic Acid | 36.5 seconds | 40023050 |
Predicted Kovats Retention IndicesUnderivatizedDerivatized |
|---|
| Spectra |
|---|
| |
|---|
| Biological Properties |
|---|
| Cellular Locations | - Cytoplasm
- Extracellular
- Membrane (predicted from logP)
- Endoplasmic reticulum
- Peroxisome
|
|---|
| Biospecimen Locations | - Blood
- Cerebrospinal Fluid (CSF)
- Feces
- Saliva
- Sweat
- Urine
|
|---|
| Tissue Locations | - Adipose Tissue
- Bladder
- Epidermis
- Fibroblasts
- Kidney
- Placenta
- Platelet
- Prostate
- Skeletal Muscle
|
|---|
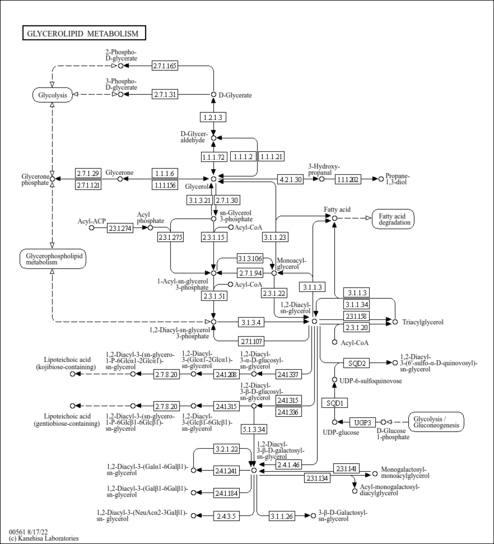
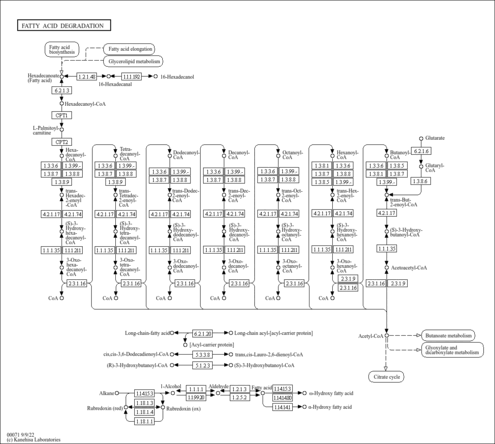
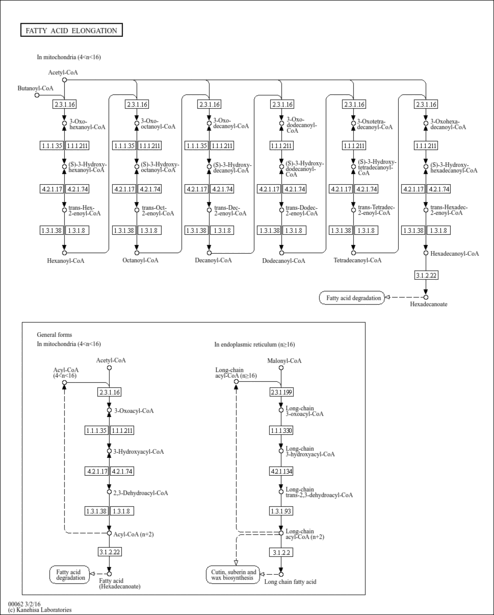
| Pathways | |
|---|
| Normal Concentrations |
|---|
| |
| Blood | Detected and Quantified | 129 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 1060.743 +/- 35.0981 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 1064.643 +/- 50.697 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 1052.943 +/- 54.597 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 1029.544 +/- 62.397 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 52 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 53 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 74 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 80 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 95 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 114 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 131 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 147 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 347 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 366 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 424 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 1415.468 +/- 323.410 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 1215.798 +/- 212.0316 uM | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected and Quantified | 2500 +/- 630 uM | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected and Quantified | 2360 +/- 430 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 22.3-31.1 uM | Adult (>18 years old) | Male | Normal | | details | | Blood | Detected and Quantified | 22.7-32.9 uM | Adult (>18 years old) | Female | Normal | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 25.0 +/- 2.8 uM | Children (1-13 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 25.5 +/- 2.8 uM | Adolescent (13-18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 26.7 +/- 4.4 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 30.49 +/- 2.64 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 66.010 +/- 9.879 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 63.8 +/- 0.400 uM | Adult (>18 years old) | Both | Normal | | details | | Blood | Detected and Quantified | 122 +/- 48 uM | Adult (>18 years old) | Both | Normal | | details | | Cerebrospinal Fluid (CSF) | Detected and Quantified | 18.0 +/- 12.0 uM | Adult (>18 years old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected and Quantified | 122.219 +/- 200.254 nmol/g wet feces | Not Specified | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Infant (0-1 year old) | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Not Specified | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Not Specified | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Children (1-13 years old) | Both | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Infant (0-1 year old) | Not Available | Normal | | details | | Feces | Detected and Quantified | 421.957 +/- 602.517 nmol/g wet feces | Not Specified | Not Specified | Normal | | details | | Feces | Detected but not Quantified | Not Quantified | Infant (0-1 year old) | Not Specified | Normal | | details | | Saliva | Detected and Quantified | 11.09 +/- 2.5 uM | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Not Specified | Normal | | details | | Saliva | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Male | Normal | | details | | Sweat | Detected but not Quantified | Not Quantified | Adult | Both | Normal | | details | | Urine | Detected and Quantified | 3.5-7.2 umol/mmol creatinine | Infant (0-1 year old) | Female | Normal | | details | | Urine | Detected and Quantified | 3.0-8.7 umol/mmol creatinine | Infant (0-1 year old) | Male | Normal | | details | | Urine | Detected and Quantified | 4-10.5 umol/mmol creatinine | Infant (0-1 year old) | Female | Normal | | details | | Urine | Detected and Quantified | 1.4 (0.1-3.8) umol/mmol creatinine | Children (1-13 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 4.0-9.3 umol/mmol creatinine | Infant (0-1 year old) | Male | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 10.8 (2.6-32.7) umol/mmol creatinine | Newborn (0-30 days old) | Both | Normal | | details | | Urine | Detected and Quantified | 2.2 (0.8-8.2) umol/mmol creatinine | Children (1-13 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 1.4 (0.1-7.3) umol/mmol creatinine | Adolescent (13-18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.30 +/- 0.09 umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 11 (6.0-23) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 6.7(2.6-24.3) umol/mmol creatinine | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Normal | | details | | Urine | Detected and Quantified | 11.2 (6.0-26.1) umol/mmol creatinine | Infant (0-1 year old) | Both | Normal | | details | | Urine | Detected and Quantified | 0.45-0.73 umol/mmol creatinine | Adult (>18 years old) | Male | Normal | | details | | Urine | Detected and Quantified | 0.46-1.1 umol/mmol creatinine | Adult (>18 years old) | Female | Normal | | details | | Urine | Detected and Quantified | 16.903 +/- 12.225 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Normal | | details |
|
|---|
| Abnormal Concentrations |
|---|
| |
| Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Female | Breast cancer | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Schizophrenia | | details | | Blood | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Schizophrenia | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | CCD | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Ileal Crohn's disease | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Inflammatory bowel disease | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal Cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Newborn (0-30 days old) | Not Specified | Premature neonates | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Feces | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Colorectal cancer | | details | | Urine | Detected and Quantified | 22.852 +/- 19.074 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Eosinophilic esophagitis | | details | | Urine | Detected and Quantified | 10.703 +/- 11.558 umol/mmol creatinine | Children (1 - 13 years old) | Not Specified | Gastroesophageal reflux disease | | details | | Urine | Detected but not Quantified | Not Quantified | Adult (>18 years old) | Both | Bladder cancer | | details |
|
|---|
| Associated Disorders and Diseases |
|---|
| Disease References | | Perillyl alcohol administration for cancer treatment |
|---|
- Lv W, Yang T: Identification of possible biomarkers for breast cancer from free fatty acid profiles determined by GC-MS and multivariate statistical analysis. Clin Biochem. 2012 Jan;45(1-2):127-33. doi: 10.1016/j.clinbiochem.2011.10.011. Epub 2011 Oct 26. [PubMed:22061338 ]
| | Colorectal cancer |
|---|
- Ni Y, Xie G, Jia W: Metabonomics of human colorectal cancer: new approaches for early diagnosis and biomarker discovery. J Proteome Res. 2014 Sep 5;13(9):3857-70. doi: 10.1021/pr500443c. Epub 2014 Aug 14. [PubMed:25105552 ]
- Brown DG, Rao S, Weir TL, O'Malia J, Bazan M, Brown RJ, Ryan EP: Metabolomics and metabolic pathway networks from human colorectal cancers, adjacent mucosa, and stool. Cancer Metab. 2016 Jun 6;4:11. doi: 10.1186/s40170-016-0151-y. eCollection 2016. [PubMed:27275383 ]
- Sinha R, Ahn J, Sampson JN, Shi J, Yu G, Xiong X, Hayes RB, Goedert JJ: Fecal Microbiota, Fecal Metabolome, and Colorectal Cancer Interrelations. PLoS One. 2016 Mar 25;11(3):e0152126. doi: 10.1371/journal.pone.0152126. eCollection 2016. [PubMed:27015276 ]
- Goedert JJ, Sampson JN, Moore SC, Xiao Q, Xiong X, Hayes RB, Ahn J, Shi J, Sinha R: Fecal metabolomics: assay performance and association with colorectal cancer. Carcinogenesis. 2014 Sep;35(9):2089-96. doi: 10.1093/carcin/bgu131. Epub 2014 Jul 18. [PubMed:25037050 ]
| | Schizophrenia |
|---|
- Yang J, Chen T, Sun L, Zhao Z, Qi X, Zhou K, Cao Y, Wang X, Qiu Y, Su M, Zhao A, Wang P, Yang P, Wu J, Feng G, He L, Jia W, Wan C: Potential metabolite markers of schizophrenia. Mol Psychiatry. 2013 Jan;18(1):67-78. doi: 10.1038/mp.2011.131. Epub 2011 Oct 25. [PubMed:22024767 ]
- Xuan J, Pan G, Qiu Y, Yang L, Su M, Liu Y, Chen J, Feng G, Fang Y, Jia W, Xing Q, He L: Metabolomic profiling to identify potential serum biomarkers for schizophrenia and risperidone action. J Proteome Res. 2011 Dec 2;10(12):5433-43. doi: 10.1021/pr2006796. Epub 2011 Nov 8. [PubMed:22007635 ]
| | Inflammatory bowel disease |
|---|
- Lee T, Clavel T, Smirnov K, Schmidt A, Lagkouvardos I, Walker A, Lucio M, Michalke B, Schmitt-Kopplin P, Fedorak R, Haller D: Oral versus intravenous iron replacement therapy distinctly alters the gut microbiota and metabolome in patients with IBD. Gut. 2017 May;66(5):863-871. doi: 10.1136/gutjnl-2015-309940. Epub 2016 Feb 4. [PubMed:26848182 ]
| | Eosinophilic esophagitis |
|---|
- Slae, M., Huynh, H., Wishart, D.S. (2014). Analysis of 30 normal pediatric urine samples via NMR spectroscopy (unpublished work). NA.
|
|
|---|
| Associated OMIM IDs | |
|---|
| External Links |
|---|
| DrugBank ID | DB03796 |
|---|
| Phenol Explorer Compound ID | Not Available |
|---|
| FooDB ID | FDB011679 |
|---|
| KNApSAcK ID | C00030479 |
|---|
| Chemspider ID | 960 |
|---|
| KEGG Compound ID | C00249 |
|---|
| BioCyc ID | PALMITATE |
|---|
| BiGG ID | 34386 |
|---|
| Wikipedia Link | Palmitic_acid |
|---|
| METLIN ID | 187 |
|---|
| PubChem Compound | 985 |
|---|
| PDB ID | Not Available |
|---|
| ChEBI ID | 15756 |
|---|
| Food Biomarker Ontology | Not Available |
|---|
| VMH ID | HDCA |
|---|
| MarkerDB ID | MDB00000104 |
|---|
| Good Scents ID | rw1009101 |
|---|
| References |
|---|
| Synthesis Reference | Xu, Yan; Ling, Li. A method for preparing conjugated linoleic acid and palmitic acid. Faming Zhuanli Shenqing Gongkai Shuomingshu (2005), 5 pp. |
|---|
| Material Safety Data Sheet (MSDS) | Download (PDF) |
|---|
| General References | - Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [PubMed:19212411 ]
- Guneral F, Bachmann C: Age-related reference values for urinary organic acids in a healthy Turkish pediatric population. Clin Chem. 1994 Jun;40(6):862-6. [PubMed:8087979 ]
- Hoffmann GF, Meier-Augenstein W, Stockler S, Surtees R, Rating D, Nyhan WL: Physiology and pathophysiology of organic acids in cerebrospinal fluid. J Inherit Metab Dis. 1993;16(4):648-69. [PubMed:8412012 ]
- A J, Trygg J, Gullberg J, Johansson AI, Jonsson P, Antti H, Marklund SL, Moritz T: Extraction and GC/MS analysis of the human blood plasma metabolome. Anal Chem. 2005 Dec 15;77(24):8086-94. [PubMed:16351159 ]
- Cater NB, Denke MA: Behenic acid is a cholesterol-raising saturated fatty acid in humans. Am J Clin Nutr. 2001 Jan;73(1):41-4. [PubMed:11124748 ]
- Lloyd B, Halter RJ, Kuchan MJ, Baggs GE, Ryan AS, Masor ML: Formula tolerance in postbreastfed and exclusively formula-fed infants. Pediatrics. 1999 Jan;103(1):E7. [PubMed:9917487 ]
- Katsuta Y, Iida T, Inomata S, Denda M: Unsaturated fatty acids induce calcium influx into keratinocytes and cause abnormal differentiation of epidermis. J Invest Dermatol. 2005 May;124(5):1008-13. [PubMed:15854043 ]
- Mensink RP: Nutrition in lipid disorders. Ther Umsch. 1995 Aug;52(8):509-14. [PubMed:7676394 ]
- Mesiha MS, Ponnapula S, Plakogiannis F: Oral absorption of insulin encapsulated in artificial chyles of bile salts, palmitic acid and alpha-tocopherol dispersions. Int J Pharm. 2002 Dec 5;249(1-2):1-5. [PubMed:12433429 ]
- Dubinin DM, Naidina VP, Zaloguev SN: [Evaluation of human skin function in a sealed room by a chromatographic method]. Kosm Biol Aviakosm Med. 1985 Nov-Dec;19(6):69-73. [PubMed:2868151 ]
- Okun JG, Kolker S, Schulze A, Kohlmuller D, Olgemoller K, Lindner M, Hoffmann GF, Wanders RJ, Mayatepek E: A method for quantitative acylcarnitine profiling in human skin fibroblasts using unlabelled palmitic acid: diagnosis of fatty acid oxidation disorders and differentiation between biochemical phenotypes of MCAD deficiency. Biochim Biophys Acta. 2002 Oct 10;1584(2-3):91-8. [PubMed:12385891 ]
- Otsuki T, Oku M: [A study on the effect of cortisol and progesterone on cytosolic arachidonic and palmitic acid concentrations in cultured human myometrial cells]. Nihon Sanka Fujinka Gakkai Zasshi. 1995 Jun;47(6):531-8. [PubMed:7608616 ]
- Draisey TF, Gagneja GL, Thibert RJ: Pulmonary surfactant and amniotic fluid insulin. Obstet Gynecol. 1977 Aug;50(2):197-9. [PubMed:577607 ]
- Florentin E, Athias A, Lagrost L: Modulation of the activity of the human cholesteryl ester transfer protein by carboxylated derivatives. Evidence for 13-cis-retinoic acid as a potent activator of the protein's activity in plasma. Eur J Biochem. 1996 Sep 15;240(3):699-706. [PubMed:8856073 ]
- Urien S, Morin D, Tillement JP: Effect of alpha-1-acid glycoprotein, albumin and palmitic acid on the brain and salivary gland extraction of warfarin in rats. J Pharmacol Exp Ther. 1989 Feb;248(2):781-5. [PubMed:2918479 ]
- Wang J, Zhu F, Pan G: [Diagnosis of chylous ascites with oral administration of 13C-palmitic acid]. Zhonghua Nei Ke Za Zhi. 1996 Jun;35(6):382-4. [PubMed:9387625 ]
- Ip MP, Draisey TF, Thibert RJ, Gagneja GL, Jasey GM: Fetal lung maturity, as assessed by gas-liquid chromatographic determination of phospholipid palmitic acid in amniotic fluid. Clin Chem. 1977 Jan;23(1):35-40. [PubMed:832370 ]
- Messner T, Erkstam UB, Gustafsson IB, Nilsson SB, Vessby B: Diet and dietary markers in Kiruna and Uppsala, Sweden--a comparison. Int J Circumpolar Health. 1997 Apr;56(1-2):21-9. [PubMed:9300843 ]
- Vessby B, Tengblad S, Lithell H: Insulin sensitivity is related to the fatty acid composition of serum lipids and skeletal muscle phospholipids in 70-year-old men. Diabetologia. 1994 Oct;37(10):1044-50. [PubMed:7851683 ]
- Liau YH, Slomiany BL, Slomiany A, Piasek A, Palmer D, Rosenthal WS: Identification of mucus glycoprotein fatty acyltransferase activity in human gastric mucosa. Digestion. 1985;32(1):57-62. [PubMed:4018446 ]
- Thiele I, Swainston N, Fleming RM, Hoppe A, Sahoo S, Aurich MK, Haraldsdottir H, Mo ML, Rolfsson O, Stobbe MD, Thorleifsson SG, Agren R, Bolling C, Bordel S, Chavali AK, Dobson P, Dunn WB, Endler L, Hala D, Hucka M, Hull D, Jameson D, Jamshidi N, Jonsson JJ, Juty N, Keating S, Nookaew I, Le Novere N, Malys N, Mazein A, Papin JA, Price ND, Selkov E Sr, Sigurdsson MI, Simeonidis E, Sonnenschein N, Smallbone K, Sorokin A, van Beek JH, Weichart D, Goryanin I, Nielsen J, Westerhoff HV, Kell DB, Mendes P, Palsson BO: A community-driven global reconstruction of human metabolism. Nat Biotechnol. 2013 May;31(5):419-25. doi: 10.1038/nbt.2488. Epub 2013 Mar 3. [PubMed:23455439 ]
- KINGSBURY KJ, PAUL S, CROSSLEY A, MORGAN DM: The fatty acid composition of human depot fat. Biochem J. 1961 Mar;78:541-50. [PubMed:13756126 ]
- Jensen RG, Hagerty MM, McMahon KE: Lipids of human milk and infant formulas: a review. Am J Clin Nutr. 1978 Jun;31(6):990-1016. doi: 10.1093/ajcn/31.6.990. [PubMed:352132 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
|
|---|