| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2023-05-30 20:55:55 UTC |
|---|
| HMDB ID | HMDB0000225 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | Oxoadipic acid |
|---|
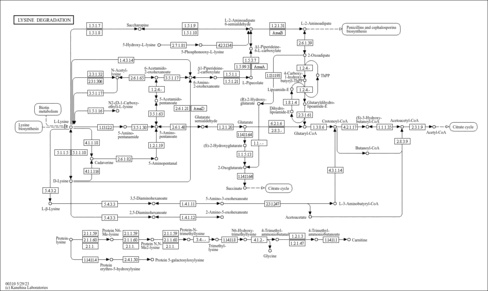
| Description | 2-Oxoadipic acid is produced from lysine in the cytosol of cells via the saccharopine and the pipecolic acid pathways. Catabolites of hydroxylysine and tryptophan enter these pathways as 2-aminoadipic- -semialdehyde and 2-oxoadipate, respectively. In the matrix of mitochondria, 2-oxoadipate is decarboxylated to glutaryl-CoA by the 2-oxoadipate dehydrogenase complex and then converted to acetyl-CoA. 2-Oxoadipic aciduria is an in-born error of metabolism of lysine, tryptophan, and hydroxylysine, in which abnormal quantities of 2-aminoadipic acid are found in body fluids along with 2-oxoadipic acid. Patients with 2-Oxoadipic acidemias are mentally retarded with hypotonia or seizures. 2-Oxoadipic aciduria can occur in patients with Kearns-Sayre Syndrome, a progressive disorder with onset prior to 20 years of age in which multiple organ systems are affected, including progressive external ophthalmoplegia, retinopathy, and the age of onset, and these are associated classically with abnormalities in cardiac conduction, cerebellar signs, and elevated cerebrospinal fluid protein (PMID: 10655159 , 16183823 , 11083877 ). Oxoadipic acid is found to be associated with alpha-aminoadipic aciduria, which is an inborn error of metabolism. |
|---|
| Structure | InChI=1S/C6H8O5/c7-4(6(10)11)2-1-3-5(8)9/h1-3H2,(H,8,9)(H,10,11) |
|---|
| Synonyms | | Value | Source |
|---|
| 2-Oxoadipate | ChEBI | | 2-Oxoadipic acid | Kegg | | Oxoadipate | Generator | | 2-Keto-adipate | HMDB | | 2-Ketoadipate | HMDB | | 2-Ketoadipic acid | HMDB | | 2-oxo-Hexanedioate | HMDB | | 2-oxo-Hexanedioic acid | HMDB | | 2-Oxohexanedioate | HMDB | | 2-Oxohexanedioic acid | HMDB | | 2-Oxohexanedionic acid | HMDB | | a-Ketoadipate | HMDB | | a-Ketoadipic acid | HMDB | | a-Oxoadipate | HMDB | | a-Oxoadipic acid | HMDB | | alpha-Ketoadipate | HMDB | | alpha-Ketoadipic acid | HMDB | | alpha-Oxoadipate | HMDB | | alpha-Oxoadipic acid | HMDB | | α-Ketoadipic acid | HMDB | | α-Oxoadipic acid | HMDB |
|
|---|
| Chemical Formula | C6H8O5 |
|---|
| Average Molecular Weight | 160.1247 |
|---|
| Monoisotopic Molecular Weight | 160.037173366 |
|---|
| IUPAC Name | 2-oxohexanedioic acid |
|---|
| Traditional Name | oxoadipate |
|---|
| CAS Registry Number | 3184-35-8 |
|---|
| SMILES | OC(=O)CCCC(=O)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C6H8O5/c7-4(6(10)11)2-1-3-5(8)9/h1-3H2,(H,8,9)(H,10,11) |
|---|
| InChI Key | FGSBNBBHOZHUBO-UHFFFAOYSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as medium-chain keto acids and derivatives. These are keto acids with a 6 to 12 carbon atoms long side chain. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Keto acids and derivatives |
|---|
| Sub Class | Medium-chain keto acids and derivatives |
|---|
| Direct Parent | Medium-chain keto acids and derivatives |
|---|
| Alternative Parents | |
|---|
| Substituents | - Medium-chain keto acid
- Dicarboxylic acid or derivatives
- Alpha-keto acid
- Alpha-hydroxy ketone
- Ketone
- Carboxylic acid
- Carboxylic acid derivative
- Organic oxygen compound
- Organic oxide
- Hydrocarbon derivative
- Organooxygen compound
- Carbonyl group
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Physiological effect | |
|---|
| Disposition | |
|---|
| Process | |
|---|
| Role | |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | 127 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | Not Available | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Experimental Collision Cross Sections |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Retention Times Underivatized| Chromatographic Method | Retention Time | Reference |
|---|
| Measured using a Waters Acquity ultraperformance liquid chromatography (UPLC) ethylene-bridged hybrid (BEH) C18 column (100 mm × 2.1 mm; 1.7 μmparticle diameter). Predicted by Afia on May 17, 2022. Predicted by Afia on May 17, 2022. | 2.39 minutes | 32390414 | | Predicted by Siyang on May 30, 2022 | 9.496 minutes | 33406817 | | Predicted by Siyang using ReTip algorithm on June 8, 2022 | 3.57 minutes | 32390414 | | AjsUoB = Accucore 150 Amide HILIC with 10mM Ammonium Formate, 0.1% Formic Acid | 163.5 seconds | 40023050 | | Fem_Long = Waters ACQUITY UPLC HSS T3 C18 with Water:MeOH and 0.1% Formic Acid | 1071.6 seconds | 40023050 | | Fem_Lipids = Ascentis Express C18 with (60:40 water:ACN):(90:10 IPA:ACN) and 10mM NH4COOH + 0.1% Formic Acid | 305.9 seconds | 40023050 | | Life_Old = Waters ACQUITY UPLC BEH C18 with Water:(20:80 acetone:ACN) and 0.1% Formic Acid | 80.6 seconds | 40023050 | | Life_New = RP Waters ACQUITY UPLC HSS T3 C18 with Water:(30:70 MeOH:ACN) and 0.1% Formic Acid | 182.1 seconds | 40023050 | | RIKEN = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 73.9 seconds | 40023050 | | Eawag_XBridgeC18 = XBridge C18 3.5u 2.1x50 mm with Water:MeOH and 0.1% Formic Acid | 247.1 seconds | 40023050 | | BfG_NTS_RP1 =Agilent Zorbax Eclipse Plus C18 (2.1 mm x 150 mm, 3.5 um) with Water:ACN and 0.1% Formic Acid | 299.0 seconds | 40023050 | | HILIC_BDD_2 = Merck SeQuant ZIC-HILIC with ACN(0.1% formic acid):water(16 mM ammonium formate) | 292.8 seconds | 40023050 | | UniToyama_Atlantis = RP Waters Atlantis T3 (2.1 x 150 mm, 5 um) with ACN:Water and 0.1% Formic Acid | 641.5 seconds | 40023050 | | BDD_C18 = Hypersil Gold 1.9µm C18 with Water:ACN and 0.1% Formic Acid | 188.2 seconds | 40023050 | | UFZ_Phenomenex = Kinetex Core-Shell C18 2.6 um, 3.0 x 100 mm, Phenomenex with Water:MeOH and 0.1% Formic Acid | 858.4 seconds | 40023050 | | SNU_RIKEN_POS = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 197.5 seconds | 40023050 | | RPMMFDA = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 229.4 seconds | 40023050 | | MTBLS87 = Merck SeQuant ZIC-pHILIC column with ACN:Water and :ammonium carbonate | 657.5 seconds | 40023050 | | KI_GIAR_zic_HILIC_pH2_7 = Merck SeQuant ZIC-HILIC with ACN:Water and 0.1% FA | 267.4 seconds | 40023050 | | Meister zic-pHILIC pH9.3 = Merck SeQuant ZIC-pHILIC column with ACN:Water 5mM NH4Ac pH9.3 and 5mM ammonium acetate in water | 389.2 seconds | 40023050 |
Predicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| Oxoadipic acid,1TMS,isomer #1 | C[Si](C)(C)OC(=O)CCCC(=O)C(=O)O | 1524.7 | Semi standard non polar | 33892256 | | Oxoadipic acid,1TMS,isomer #2 | C[Si](C)(C)OC(=O)C(=O)CCCC(=O)O | 1505.1 | Semi standard non polar | 33892256 | | Oxoadipic acid,1TMS,isomer #3 | C[Si](C)(C)OC(=CCCC(=O)O)C(=O)O | 1638.8 | Semi standard non polar | 33892256 | | Oxoadipic acid,2TMS,isomer #1 | C[Si](C)(C)OC(=O)CCCC(=O)C(=O)O[Si](C)(C)C | 1633.2 | Semi standard non polar | 33892256 | | Oxoadipic acid,2TMS,isomer #2 | C[Si](C)(C)OC(=O)CCC=C(O[Si](C)(C)C)C(=O)O | 1707.0 | Semi standard non polar | 33892256 | | Oxoadipic acid,2TMS,isomer #3 | C[Si](C)(C)OC(=O)C(=CCCC(=O)O)O[Si](C)(C)C | 1667.6 | Semi standard non polar | 33892256 | | Oxoadipic acid,3TMS,isomer #1 | C[Si](C)(C)OC(=O)CCC=C(O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1724.0 | Semi standard non polar | 33892256 | | Oxoadipic acid,3TMS,isomer #1 | C[Si](C)(C)OC(=O)CCC=C(O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1667.6 | Standard non polar | 33892256 | | Oxoadipic acid,3TMS,isomer #1 | C[Si](C)(C)OC(=O)CCC=C(O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1688.2 | Standard polar | 33892256 | | Oxoadipic acid,1TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCCC(=O)C(=O)O | 1791.4 | Semi standard non polar | 33892256 | | Oxoadipic acid,1TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)C(=O)CCCC(=O)O | 1781.1 | Semi standard non polar | 33892256 | | Oxoadipic acid,1TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=CCCC(=O)O)C(=O)O | 1881.1 | Semi standard non polar | 33892256 | | Oxoadipic acid,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCCC(=O)C(=O)O[Si](C)(C)C(C)(C)C | 2081.3 | Semi standard non polar | 33892256 | | Oxoadipic acid,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)CCC=C(O[Si](C)(C)C(C)(C)C)C(=O)O | 2163.1 | Semi standard non polar | 33892256 | | Oxoadipic acid,2TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=O)C(=CCCC(=O)O)O[Si](C)(C)C(C)(C)C | 2132.1 | Semi standard non polar | 33892256 | | Oxoadipic acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCC=C(O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2356.7 | Semi standard non polar | 33892256 | | Oxoadipic acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCC=C(O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2206.4 | Standard non polar | 33892256 | | Oxoadipic acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCC=C(O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2123.7 | Standard polar | 33892256 |
|
|---|
| GC-MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental GC-MS | GC-MS Spectrum - Oxoadipic acid GC-EI-TOF (Non-derivatized) | splash10-002b-3910000000-66f39a8b247786a425e6 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoadipic acid GC-EI-TOF (Non-derivatized) | splash10-002k-3910000000-977efef811bb47535b80 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoadipic acid GC-EI-TOF (Non-derivatized) | splash10-0udr-6900000000-0eb27647b831277a2aef | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoadipic acid GC-EI-TOF (Non-derivatized) | splash10-0udr-6900000000-46cd4d782e3d19030993 | 2017-09-12 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoadipic acid GC-EI-TOF (Non-derivatized) | splash10-002b-3910000000-66f39a8b247786a425e6 | 2018-05-18 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoadipic acid GC-EI-TOF (Non-derivatized) | splash10-002k-3910000000-977efef811bb47535b80 | 2018-05-18 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoadipic acid GC-EI-TOF (Non-derivatized) | splash10-0udr-6900000000-0eb27647b831277a2aef | 2018-05-18 | HMDB team, MONA, MassBank | View Spectrum | | Experimental GC-MS | GC-MS Spectrum - Oxoadipic acid GC-EI-TOF (Non-derivatized) | splash10-0udr-6900000000-46cd4d782e3d19030993 | 2018-05-18 | HMDB team, MONA, MassBank | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (Non-derivatized) - 70eV, Positive | splash10-00r6-9300000000-b3f156c173adabf3f1bb | 2016-09-22 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (2 TMS) - 70eV, Positive | splash10-0079-9830000000-a3b1d5e6d378582a6fc2 | 2017-10-06 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (Non-derivatized) - 70eV, Positive | Not Available | 2021-10-12 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (TMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (TMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (TMS_1_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (TMS_2_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (TMS_2_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (TBDMS_1_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (TBDMS_1_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (TBDMS_1_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (TBDMS_2_1) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (TBDMS_2_2) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum | | Predicted GC-MS | Predicted GC-MS Spectrum - Oxoadipic acid GC-MS (TBDMS_2_3) - 70eV, Positive | Not Available | 2021-11-05 | Wishart Lab | View Spectrum |
MS/MS Spectra| Spectrum Type | Description | Splash Key | Deposition Date | Source | View |
|---|
| Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid LC-ESI-QQ , negative-QTOF | splash10-0a4i-0900000000-aef94cc19e461712a097 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid LC-ESI-QQ , negative-QTOF | splash10-0bt9-9500000000-9326906313591bb9da1c | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid LC-ESI-QQ , negative-QTOF | splash10-0a4i-9000000000-05db676896bb1e350497 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid LC-ESI-QQ , negative-QTOF | splash10-0a4l-9000000000-1e566f572b14692461f2 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid LC-ESI-QQ , negative-QTOF | splash10-0006-9000000000-527da1c1c271db4d8db4 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid , negative-QTOF | splash10-0a4i-9600000000-38ade0e080ecb7a89588 | 2017-09-14 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid 20V, Negative-QTOF | splash10-0a4i-9000000000-ea231700b469554fcdb4 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid 10V, Negative-QTOF | splash10-0a4i-9300000000-d69c9552085fc7db91fc | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid 30V, Negative-QTOF | splash10-052f-9000000000-fa207039b6913600e1b4 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid 40V, Negative-QTOF | splash10-0006-9000000000-dc580b3d8a9e58ea79c6 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid 10V, Negative-QTOF | splash10-0a4i-9200000000-f25470d83f4df952958e | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid 20V, Negative-QTOF | splash10-0a4i-9000000000-c524dfcde428d896e041 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Experimental LC-MS/MS | LC-MS/MS Spectrum - Oxoadipic acid 40V, Negative-QTOF | splash10-0006-9000000000-3e6aec4efdb02abcf887 | 2021-09-20 | HMDB team, MONA | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 10V, Positive-QTOF | splash10-0006-2900000000-757eb57780db9df4f78d | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 20V, Positive-QTOF | splash10-016v-9700000000-675e5732632f0a0cabe1 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 40V, Positive-QTOF | splash10-0006-9000000000-1da8154067fdb9a27459 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 10V, Negative-QTOF | splash10-0a4i-1900000000-0e26a52af95eb287809d | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 20V, Negative-QTOF | splash10-066v-7900000000-d898e0593464c13b3f89 | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 40V, Negative-QTOF | splash10-052g-9100000000-34782bad1c8755ac897d | 2015-09-15 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 10V, Positive-QTOF | splash10-014v-9800000000-1b44ce4709a42734f384 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 20V, Positive-QTOF | splash10-00kf-9000000000-e16740a0d326b489193c | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 40V, Positive-QTOF | splash10-0006-9000000000-99207abec9ffe380ff34 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 10V, Negative-QTOF | splash10-0295-5900000000-d4320b8980ff5d5a03d3 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 20V, Negative-QTOF | splash10-00kb-9100000000-27e2c4cbbb4945978dc1 | 2021-09-24 | Wishart Lab | View Spectrum | | Predicted LC-MS/MS | Predicted LC-MS/MS Spectrum - Oxoadipic acid 40V, Negative-QTOF | splash10-0006-9000000000-bc42e02ad867f6ce85a4 | 2021-09-24 | Wishart Lab | View Spectrum |
NMR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Experimental 1D NMR | 1H NMR Spectrum (1D, 500 MHz, H2O, experimental) | 2012-12-04 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 100 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 1000 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 200 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 300 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 400 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 500 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 600 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 700 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 800 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 13C NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Predicted 1D NMR | 1H NMR Spectrum (1D, 900 MHz, D2O, predicted) | 2021-09-29 | Wishart Lab | View Spectrum | | Experimental 2D NMR | [1H, 13C]-HSQC NMR Spectrum (2D, 500 MHz, H2O, experimental) | 2012-12-05 | Wishart Lab | View Spectrum |
IR Spectra| Spectrum Type | Description | Deposition Date | Source | View |
|---|
| Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M-H]-) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+H]+) | 2023-02-03 | FELIX lab | View Spectrum | | Predicted IR Spectrum | IR Ion Spectrum (Predicted IRIS Spectrum, Adduct: [M+Na]+) | 2023-02-03 | FELIX lab | View Spectrum |
|
|---|
| Disease References | | 2-Ketoadipic acidemia |
|---|
- Chalmers RA, Lawson AM, Watts RW, Tavill AS, Kamerling JP, Hey E, Ogilvie D: D-2-hydroxyglutaric aciduria: case report and biochemical studies. J Inherit Metab Dis. 1980;3(1):11-5. [PubMed:6774165 ]
- Peng H, Shinka T, Inoue Y, Mitsubuchi H, Ishimatsu J, Yoshino M, Kuhara T: Asymptomatic alpha-ketoadipic aciduria detected during a pilot study of neonatal urine screening. Acta Paediatr. 1999 Aug;88(8):911-4. [PubMed:10503694 ]
- G.Frauendienst-Egger, Friedrich K. Trefz (2017). MetaGene: Metabolic & Genetic Information Center (MIC: http://www.metagene.de). METAGENE consortium.
| | Alpha-aminoadipic aciduria |
|---|
- Gray RG, O'Neill EM, Pollitt RJ: Alpha-aminoadipic aciduria: chemical and enzymatic studies. J Inherit Metab Dis. 1980;2(4):89-92. [PubMed:6796766 ]
| | 2-Aminoadipic aciduria |
|---|
- Danhauser K, Sauer SW, Haack TB, Wieland T, Staufner C, Graf E, Zschocke J, Strom TM, Traub T, Okun JG, Meitinger T, Hoffmann GF, Prokisch H, Kolker S: DHTKD1 mutations cause 2-aminoadipic and 2-oxoadipic aciduria. Am J Hum Genet. 2012 Dec 7;91(6):1082-7. doi: 10.1016/j.ajhg.2012.10.006. Epub 2012 Nov 8. [PubMed:23141293 ]
|
|
|---|
| General References | - Fiermonte G, Dolce V, Palmieri L, Ventura M, Runswick MJ, Palmieri F, Walker JE: Identification of the human mitochondrial oxodicarboxylate carrier. Bacterial expression, reconstitution, functional characterization, tissue distribution, and chromosomal location. J Biol Chem. 2001 Mar 16;276(11):8225-30. Epub 2000 Nov 16. [PubMed:11083877 ]
- Lee SH, Kim SO, Chung BC: Gas chromatographic-mass spectrometric determination of urinary oxoacids using O-(2,3,4,5,6-pentafluorobenzyl)oxime-trimethylsilyl ester derivatization and cation-exchange chromatography. J Chromatogr B Biomed Sci Appl. 1998 Nov 20;719(1-2):1-7. [PubMed:9869358 ]
- Guneral F, Bachmann C: Age-related reference values for urinary organic acids in a healthy Turkish pediatric population. Clin Chem. 1994 Jun;40(6):862-6. [PubMed:8087979 ]
- Barshop BA, Nyhan WL, Naviaux RK, McGowan KA, Friedlander M, Haas RH: Kearns-Sayre syndrome presenting as 2-oxoadipic aciduria. Mol Genet Metab. 2000 Jan;69(1):64-8. [PubMed:10655159 ]
- Schulze-Bergkamen A, Okun JG, Spiekerkotter U, Lindner M, Haas D, Kohlmuller D, Mayatepek E, Schulze-Bergkamen H, Greenberg CR, Zschocke J, Hoffmann GF, Kolker S: Quantitative acylcarnitine profiling in peripheral blood mononuclear cells using in vitro loading with palmitic and 2-oxoadipic acids: biochemical confirmation of fatty acid oxidation and organic acid disorders. Pediatr Res. 2005 Nov;58(5):873-80. Epub 2005 Sep 23. [PubMed:16183823 ]
|
|---|