| Record Information |
|---|
| Version | 5.0 |
|---|
| Status | Detected and Quantified |
|---|
| Creation Date | 2005-11-16 15:48:42 UTC |
|---|
| Update Date | 2025-05-29 19:32:58 UTC |
|---|
| HMDB ID | HMDB0000510 |
|---|
| Secondary Accession Numbers | |
|---|
| Metabolite Identification |
|---|
| Common Name | Aminoadipic acid |
|---|
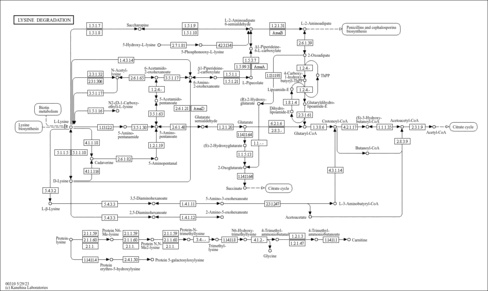
| Description | Aminoadipic acid (CAS: 542-32-5), also known as 2-aminoadipate, is a metabolite in the principal biochemical pathway of lysine. It is an intermediate in the metabolism (i.e. breakdown or degradation) of lysine and saccharopine. It antagonizes neuroexcitatory activity modulated by the glutamate receptor N-methyl-D-aspartate (NMDA). Aminoadipic acid has also been shown to inhibit the production of kynurenic acid, a broad spectrum excitatory amino acid receptor antagonist, in brain tissue slices (PMID: 8566117 ). Recent studies have shown that aminoadipic acid is elevated in prostate biopsy tissues from prostate cancer patients (PMID: 23737455 ). Mutations in DHTKD1 (dehydrogenase E1 and transketolase domain-containing protein 1) have been shown to cause human 2-aminoadipic aciduria and 2-oxoadipic aciduria via impaired decarboxylation of 2-oxoadipate to glutaryl-CoA, which is the last step in the lysine degradation pathway (PMID: 23141293 ). Aging, diabetes, sepsis, and renal failure are known to catalyze the oxidation of lysyl residues to form 2-aminoadipic acid in human skin collagen and potentially other tissues (PMID: 18448817 ). Proteolytic breakdown of these tissues can lead to the release of free 2-aminoadipic acid. Studies in rats indicate that aminoadipic acid (along with the three branched-chain amino acids: leucine, valine, and isoleucine) levels are elevated in the pre-diabetic phase and so aminoadipic acid may serve as a predictive biomarker for the development of diabetes (PMID: 15389298 ). Long-term hyperglycemia of endothelial cells can also lead to elevated levels of aminoadipate which is thought to be a sign of lysine breakdown through oxidative stress and reactive oxygen species (ROS) (PMID: 21961526 ). 2-Aminoadipate is a potential small-molecule marker of oxidative stress (PMID: 21647514 ). Therefore, depending on the circumstances aminoadipic acid can act as an acidogen, a diabetogen, an atherogen, and a metabotoxin. An acidogen is an acidic compound that induces acidosis, which has multiple adverse effects on many organ systems. A diabetogen is a compound that can lead to type 2 diabetes. An atherogen is a compound that leads to atherosclerosis and cardiovascular disease. A metabotoxin is an endogenously produced metabolite that causes adverse health effects at chronically high levels. Chronically high levels of aminoadipic acid are associated with at least two inborn errors of metabolism including 2-aminoadipic aciduria and 2-oxoadipic aciduria. Aminoadipic acid is an organic acid and abnormally high levels of organic acids in the blood (organic acidemia), urine (organic aciduria), the brain, and other tissues lead to general metabolic acidosis. Acidosis typically occurs when arterial pH falls below 7.35. In infants with acidosis, the initial symptoms include poor feeding, vomiting, loss of appetite, weak muscle tone (hypotonia), and lack of energy (lethargy). These can progress to heart abnormalities, kidney abnormalities, liver damage, seizures, coma, and possibly death. These are also the characteristic symptoms of the untreated IEMs mentioned above. Many affected children with organic acidemias experience intellectual disability or delayed development. In adults, acidosis or acidemia is characterized by headaches, confusion, feeling tired, tremors, sleepiness, and seizures. As a diabetogen, serum aminoadipic levels appear to regulate glucose homeostasis and have been highly predictive of individuals who later develop diabetes (PMID: 24091325 ). In particular, aminoadipic acid lowers fasting plasma glucose levels and enhances insulin secretion from human islets. As an atherogen, aminoadipic acid has been found to be produced at high levels via protein lysine oxidation in atherosclerotic plaques (PMID: 28069522 ). |
|---|
| Structure | InChI=1S/C6H11NO4/c7-4(6(10)11)2-1-3-5(8)9/h4H,1-3,7H2,(H,8,9)(H,10,11)/t4-/m0/s1 |
|---|
| Synonyms | | Value | Source |
|---|
| (S)-2-Aminohexanedioic acid | ChEBI | | L-2-Aminoadipate | ChEBI | | L-2-Aminohexanedioate | ChEBI | | L-alpha-Aminoadipate | ChEBI | | L-alpha-Aminoadipic acid | ChEBI | | L-2-Aminoadipic acid | Kegg | | (S)-2-Aminohexanedioate | Generator | | L-2-Aminohexanedioic acid | Generator | | L-a-Aminoadipate | Generator | | L-a-Aminoadipic acid | Generator | | L-Α-aminoadipate | Generator | | L-Α-aminoadipic acid | Generator | | Aminoadipate | Generator | | (2S)-2-Aminohexanedioic acid | HMDB | | (2S)-2-Azaniumyl-6-hydroxy-6-oxohexanoate | HMDB | | (S)-2-Aminoadipic acid | HMDB | | 2-Aminoadipate | HMDB | | 2-Aminoadipic acid | HMDB | | 2-Aminohexanedioic acid | HMDB | | L-Aminoadipic acid | HMDB | | alpha-Aminoadipic acid | HMDB | | Α-aminoadipic acid | HMDB |
|
|---|
| Chemical Formula | C6H11NO4 |
|---|
| Average Molecular Weight | 161.1558 |
|---|
| Monoisotopic Molecular Weight | 161.068807845 |
|---|
| IUPAC Name | (2S)-2-aminohexanedioic acid |
|---|
| Traditional Name | aminoadipate |
|---|
| CAS Registry Number | 1118-90-7 |
|---|
| SMILES | N[C@@H](CCCC(O)=O)C(O)=O |
|---|
| InChI Identifier | InChI=1S/C6H11NO4/c7-4(6(10)11)2-1-3-5(8)9/h4H,1-3,7H2,(H,8,9)(H,10,11)/t4-/m0/s1 |
|---|
| InChI Key | OYIFNHCXNCRBQI-BYPYZUCNSA-N |
|---|
| Chemical Taxonomy |
|---|
| Description | Belongs to the class of organic compounds known as l-alpha-amino acids. These are alpha amino acids which have the L-configuration of the alpha-carbon atom. |
|---|
| Kingdom | Organic compounds |
|---|
| Super Class | Organic acids and derivatives |
|---|
| Class | Carboxylic acids and derivatives |
|---|
| Sub Class | Amino acids, peptides, and analogues |
|---|
| Direct Parent | L-alpha-amino acids |
|---|
| Alternative Parents | |
|---|
| Substituents | - L-alpha-amino acid
- Medium-chain fatty acid
- Amino fatty acid
- Dicarboxylic acid or derivatives
- Fatty acid
- Fatty acyl
- Amino acid
- Carboxylic acid
- Organopnictogen compound
- Organic nitrogen compound
- Amine
- Organic oxygen compound
- Primary amine
- Organooxygen compound
- Organonitrogen compound
- Primary aliphatic amine
- Carbonyl group
- Hydrocarbon derivative
- Organic oxide
- Aliphatic acyclic compound
|
|---|
| Molecular Framework | Aliphatic acyclic compounds |
|---|
| External Descriptors | |
|---|
| Ontology |
|---|
| Not Available | Not Available |
|---|
| Physical Properties |
|---|
| State | Solid |
|---|
| Experimental Molecular Properties | | Property | Value | Reference |
|---|
| Melting Point | 196 - 198 °C | Not Available | | Boiling Point | Not Available | Not Available | | Water Solubility | 2.2 mg/mL | Not Available | | LogP | Not Available | Not Available |
|
|---|
| Experimental Chromatographic Properties | Not Available |
|---|
| Predicted Molecular Properties | |
|---|
| Predicted Chromatographic Properties | Predicted Collision Cross SectionsPredicted Retention Times Underivatized| Chromatographic Method | Retention Time | Reference |
|---|
| Measured using a Waters Acquity ultraperformance liquid chromatography (UPLC) ethylene-bridged hybrid (BEH) C18 column (100 mm × 2.1 mm; 1.7 μmparticle diameter). Predicted by Afia on May 17, 2022. Predicted by Afia on May 17, 2022. | 0.97 minutes | 32390414 | | Predicted by Siyang on May 30, 2022 | 9.3472 minutes | 33406817 | | Predicted by Siyang using ReTip algorithm on June 8, 2022 | 8.04 minutes | 32390414 | | AjsUoB = Accucore 150 Amide HILIC with 10mM Ammonium Formate, 0.1% Formic Acid | 399.8 seconds | 40023050 | | Fem_Long = Waters ACQUITY UPLC HSS T3 C18 with Water:MeOH and 0.1% Formic Acid | 511.0 seconds | 40023050 | | Fem_Lipids = Ascentis Express C18 with (60:40 water:ACN):(90:10 IPA:ACN) and 10mM NH4COOH + 0.1% Formic Acid | 299.3 seconds | 40023050 | | Life_Old = Waters ACQUITY UPLC BEH C18 with Water:(20:80 acetone:ACN) and 0.1% Formic Acid | 48.6 seconds | 40023050 | | Life_New = RP Waters ACQUITY UPLC HSS T3 C18 with Water:(30:70 MeOH:ACN) and 0.1% Formic Acid | 176.4 seconds | 40023050 | | RIKEN = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 62.6 seconds | 40023050 | | Eawag_XBridgeC18 = XBridge C18 3.5u 2.1x50 mm with Water:MeOH and 0.1% Formic Acid | 271.4 seconds | 40023050 | | BfG_NTS_RP1 =Agilent Zorbax Eclipse Plus C18 (2.1 mm x 150 mm, 3.5 um) with Water:ACN and 0.1% Formic Acid | 225.5 seconds | 40023050 | | HILIC_BDD_2 = Merck SeQuant ZIC-HILIC with ACN(0.1% formic acid):water(16 mM ammonium formate) | 811.1 seconds | 40023050 | | UniToyama_Atlantis = RP Waters Atlantis T3 (2.1 x 150 mm, 5 um) with ACN:Water and 0.1% Formic Acid | 593.2 seconds | 40023050 | | BDD_C18 = Hypersil Gold 1.9µm C18 with Water:ACN and 0.1% Formic Acid | 37.8 seconds | 40023050 | | UFZ_Phenomenex = Kinetex Core-Shell C18 2.6 um, 3.0 x 100 mm, Phenomenex with Water:MeOH and 0.1% Formic Acid | 675.4 seconds | 40023050 | | SNU_RIKEN_POS = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 175.7 seconds | 40023050 | | RPMMFDA = Waters ACQUITY UPLC BEH C18 with Water:ACN and 0.1% Formic Acid | 226.4 seconds | 40023050 | | MTBLS87 = Merck SeQuant ZIC-pHILIC column with ACN:Water and :ammonium carbonate | 806.6 seconds | 40023050 | | KI_GIAR_zic_HILIC_pH2_7 = Merck SeQuant ZIC-HILIC with ACN:Water and 0.1% FA | 490.6 seconds | 40023050 | | Meister zic-pHILIC pH9.3 = Merck SeQuant ZIC-pHILIC column with ACN:Water 5mM NH4Ac pH9.3 and 5mM ammonium acetate in water | 478.3 seconds | 40023050 |
Predicted Kovats Retention IndicesUnderivatizedDerivatized| Derivative Name / Structure | SMILES | Kovats RI Value | Column Type | Reference |
|---|
| Aminoadipic acid,1TMS,isomer #1 | C[Si](C)(C)OC(=O)CCC[C@H](N)C(=O)O | 1613.1 | Semi standard non polar | 33892256 | | Aminoadipic acid,1TMS,isomer #2 | C[Si](C)(C)OC(=O)[C@@H](N)CCCC(=O)O | 1567.0 | Semi standard non polar | 33892256 | | Aminoadipic acid,1TMS,isomer #3 | C[Si](C)(C)N[C@@H](CCCC(=O)O)C(=O)O | 1630.7 | Semi standard non polar | 33892256 | | Aminoadipic acid,2TMS,isomer #1 | C[Si](C)(C)OC(=O)CCC[C@H](N)C(=O)O[Si](C)(C)C | 1619.8 | Semi standard non polar | 33892256 | | Aminoadipic acid,2TMS,isomer #2 | C[Si](C)(C)N[C@@H](CCCC(=O)O[Si](C)(C)C)C(=O)O | 1706.9 | Semi standard non polar | 33892256 | | Aminoadipic acid,2TMS,isomer #3 | C[Si](C)(C)N[C@@H](CCCC(=O)O)C(=O)O[Si](C)(C)C | 1676.2 | Semi standard non polar | 33892256 | | Aminoadipic acid,2TMS,isomer #4 | C[Si](C)(C)N([C@@H](CCCC(=O)O)C(=O)O)[Si](C)(C)C | 1829.2 | Semi standard non polar | 33892256 | | Aminoadipic acid,3TMS,isomer #1 | C[Si](C)(C)N[C@@H](CCCC(=O)O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1723.0 | Semi standard non polar | 33892256 | | Aminoadipic acid,3TMS,isomer #1 | C[Si](C)(C)N[C@@H](CCCC(=O)O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1763.3 | Standard non polar | 33892256 | | Aminoadipic acid,3TMS,isomer #1 | C[Si](C)(C)N[C@@H](CCCC(=O)O[Si](C)(C)C)C(=O)O[Si](C)(C)C | 1896.1 | Standard polar | 33892256 | | Aminoadipic acid,3TMS,isomer #2 | C[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1874.8 | Semi standard non polar | 33892256 | | Aminoadipic acid,3TMS,isomer #2 | C[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1807.8 | Standard non polar | 33892256 | | Aminoadipic acid,3TMS,isomer #2 | C[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 2057.4 | Standard polar | 33892256 | | Aminoadipic acid,3TMS,isomer #3 | C[Si](C)(C)OC(=O)[C@H](CCCC(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1861.9 | Semi standard non polar | 33892256 | | Aminoadipic acid,3TMS,isomer #3 | C[Si](C)(C)OC(=O)[C@H](CCCC(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 1800.2 | Standard non polar | 33892256 | | Aminoadipic acid,3TMS,isomer #3 | C[Si](C)(C)OC(=O)[C@H](CCCC(=O)O)N([Si](C)(C)C)[Si](C)(C)C | 2049.0 | Standard polar | 33892256 | | Aminoadipic acid,4TMS,isomer #1 | C[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 1903.7 | Semi standard non polar | 33892256 | | Aminoadipic acid,4TMS,isomer #1 | C[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 1865.6 | Standard non polar | 33892256 | | Aminoadipic acid,4TMS,isomer #1 | C[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O[Si](C)(C)C)N([Si](C)(C)C)[Si](C)(C)C | 1822.6 | Standard polar | 33892256 | | Aminoadipic acid,1TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCC[C@H](N)C(=O)O | 1867.3 | Semi standard non polar | 33892256 | | Aminoadipic acid,1TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)[C@@H](N)CCCC(=O)O | 1833.1 | Semi standard non polar | 33892256 | | Aminoadipic acid,1TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N[C@@H](CCCC(=O)O)C(=O)O | 1904.3 | Semi standard non polar | 33892256 | | Aminoadipic acid,2TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCC[C@H](N)C(=O)O[Si](C)(C)C(C)(C)C | 2076.3 | Semi standard non polar | 33892256 | | Aminoadipic acid,2TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)N[C@@H](CCCC(=O)O[Si](C)(C)C(C)(C)C)C(=O)O | 2204.0 | Semi standard non polar | 33892256 | | Aminoadipic acid,2TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)N[C@@H](CCCC(=O)O)C(=O)O[Si](C)(C)C(C)(C)C | 2135.3 | Semi standard non polar | 33892256 | | Aminoadipic acid,2TBDMS,isomer #4 | CC(C)(C)[Si](C)(C)N([C@@H](CCCC(=O)O)C(=O)O)[Si](C)(C)C(C)(C)C | 2280.7 | Semi standard non polar | 33892256 | | Aminoadipic acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CCCC(=O)O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2381.4 | Semi standard non polar | 33892256 | | Aminoadipic acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CCCC(=O)O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2343.2 | Standard non polar | 33892256 | | Aminoadipic acid,3TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)N[C@@H](CCCC(=O)O[Si](C)(C)C(C)(C)C)C(=O)O[Si](C)(C)C(C)(C)C | 2272.6 | Standard polar | 33892256 | | Aminoadipic acid,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2549.6 | Semi standard non polar | 33892256 | | Aminoadipic acid,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2399.4 | Standard non polar | 33892256 | | Aminoadipic acid,3TBDMS,isomer #2 | CC(C)(C)[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2342.7 | Standard polar | 33892256 | | Aminoadipic acid,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CCCC(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2526.9 | Semi standard non polar | 33892256 | | Aminoadipic acid,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CCCC(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2370.5 | Standard non polar | 33892256 | | Aminoadipic acid,3TBDMS,isomer #3 | CC(C)(C)[Si](C)(C)OC(=O)[C@H](CCCC(=O)O)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2341.2 | Standard polar | 33892256 | | Aminoadipic acid,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2774.1 | Semi standard non polar | 33892256 | | Aminoadipic acid,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2607.9 | Standard non polar | 33892256 | | Aminoadipic acid,4TBDMS,isomer #1 | CC(C)(C)[Si](C)(C)OC(=O)CCC[C@@H](C(=O)O[Si](C)(C)C(C)(C)C)N([Si](C)(C)C(C)(C)C)[Si](C)(C)C(C)(C)C | 2313.2 | Standard polar | 33892256 |
|
|---|
| Disease References | | Alpha-aminoadipic aciduria |
|---|
- Vallat C, Rivier F, Bellet H, Magnan de Bornier B, Mion H, Echenne B: Treatment with vigabatrin may mimic alpha-aminoadipic aciduria. Epilepsia. 1996 Aug;37(8):803-5. [PubMed:8764822 ]
- Gray RG, O'Neill EM, Pollitt RJ: Alpha-aminoadipic aciduria: chemical and enzymatic studies. J Inherit Metab Dis. 1980;2(4):89-92. [PubMed:6796766 ]
- G.Frauendienst-Egger, Friedrich K. Trefz (2017). MetaGene: Metabolic & Genetic Information Center (MIC: http://www.metagene.de). METAGENE consortium.
| | 2-Ketoadipic acidemia |
|---|
- Chalmers RA, Lawson AM, Watts RW, Tavill AS, Kamerling JP, Hey E, Ogilvie D: D-2-hydroxyglutaric aciduria: case report and biochemical studies. J Inherit Metab Dis. 1980;3(1):11-5. [PubMed:6774165 ]
- Peng H, Shinka T, Inoue Y, Mitsubuchi H, Ishimatsu J, Yoshino M, Kuhara T: Asymptomatic alpha-ketoadipic aciduria detected during a pilot study of neonatal urine screening. Acta Paediatr. 1999 Aug;88(8):911-4. [PubMed:10503694 ]
- G.Frauendienst-Egger, Friedrich K. Trefz (2017). MetaGene: Metabolic & Genetic Information Center (MIC: http://www.metagene.de). METAGENE consortium.
| | Ulcerative colitis |
|---|
- Kolho KL, Pessia A, Jaakkola T, de Vos WM, Velagapudi V: Faecal and Serum Metabolomics in Paediatric Inflammatory Bowel Disease. J Crohns Colitis. 2017 Mar 1;11(3):321-334. doi: 10.1093/ecco-jcc/jjw158. [PubMed:27609529 ]
| | Crohn's disease |
|---|
- Kolho KL, Pessia A, Jaakkola T, de Vos WM, Velagapudi V: Faecal and Serum Metabolomics in Paediatric Inflammatory Bowel Disease. J Crohns Colitis. 2017 Mar 1;11(3):321-334. doi: 10.1093/ecco-jcc/jjw158. [PubMed:27609529 ]
| | Metastatic melanoma |
|---|
- Frankel AE, Coughlin LA, Kim J, Froehlich TW, Xie Y, Frenkel EP, Koh AY: Metagenomic Shotgun Sequencing and Unbiased Metabolomic Profiling Identify Specific Human Gut Microbiota and Metabolites Associated with Immune Checkpoint Therapy Efficacy in Melanoma Patients. Neoplasia. 2017 Oct;19(10):848-855. doi: 10.1016/j.neo.2017.08.004. Epub 2017 Sep 15. [PubMed:28923537 ]
| | Colorectal cancer |
|---|
- Sinha R, Ahn J, Sampson JN, Shi J, Yu G, Xiong X, Hayes RB, Goedert JJ: Fecal Microbiota, Fecal Metabolome, and Colorectal Cancer Interrelations. PLoS One. 2016 Mar 25;11(3):e0152126. doi: 10.1371/journal.pone.0152126. eCollection 2016. [PubMed:27015276 ]
- Goedert JJ, Sampson JN, Moore SC, Xiao Q, Xiong X, Hayes RB, Ahn J, Shi J, Sinha R: Fecal metabolomics: assay performance and association with colorectal cancer. Carcinogenesis. 2014 Sep;35(9):2089-96. doi: 10.1093/carcin/bgu131. Epub 2014 Jul 18. [PubMed:25037050 ]
| | Eosinophilic esophagitis |
|---|
- Slae, M., Huynh, H., Wishart, D.S. (2014). Analysis of 30 normal pediatric urine samples via NMR spectroscopy (unpublished work). NA.
| | Schizophrenia |
|---|
- Yang J, Chen T, Sun L, Zhao Z, Qi X, Zhou K, Cao Y, Wang X, Qiu Y, Su M, Zhao A, Wang P, Yang P, Wu J, Feng G, He L, Jia W, Wan C: Potential metabolite markers of schizophrenia. Mol Psychiatry. 2013 Jan;18(1):67-78. doi: 10.1038/mp.2011.131. Epub 2011 Oct 25. [PubMed:22024767 ]
| | Alpha-aminoadipic and alpha-ketoadipic aciduria |
|---|
- Takechi T, Okada T, Wakiguchi H, Morita H, Kurashige T, Sugahara K, Kodama H: Identification of N-acetyl-alpha-aminoadipic acid in the urine of a patient with alpha-aminoadipic and alpha-ketoadipic aciduria. J Inherit Metab Dis. 1993;16(1):119-26. [PubMed:8487492 ]
|
|
|---|
| General References | - Fiermonte G, Dolce V, Palmieri L, Ventura M, Runswick MJ, Palmieri F, Walker JE: Identification of the human mitochondrial oxodicarboxylate carrier. Bacterial expression, reconstitution, functional characterization, tissue distribution, and chromosomal location. J Biol Chem. 2001 Mar 16;276(11):8225-30. Epub 2000 Nov 16. [PubMed:11083877 ]
- Sreekumar A, Poisson LM, Rajendiran TM, Khan AP, Cao Q, Yu J, Laxman B, Mehra R, Lonigro RJ, Li Y, Nyati MK, Ahsan A, Kalyana-Sundaram S, Han B, Cao X, Byun J, Omenn GS, Ghosh D, Pennathur S, Alexander DC, Berger A, Shuster JR, Wei JT, Varambally S, Beecher C, Chinnaiyan AM: Metabolomic profiles delineate potential role for sarcosine in prostate cancer progression. Nature. 2009 Feb 12;457(7231):910-4. doi: 10.1038/nature07762. [PubMed:19212411 ]
- Brauner-Osborne H, Slok FA, Skjaerbaek N, Ebert B, Sekiyama N, Nakanishi S, Krogsgaard-Larsen P: A new highly selective metabotropic excitatory amino acid agonist: 2-amino-4-(3-hydroxy-5-methylisoxazol-4-yl)butyric acid. J Med Chem. 1996 Aug 2;39(16):3188-94. [PubMed:8759641 ]
- Hori S: [Pathophysiology of intraocular neovascularization]. Nippon Ganka Gakkai Zasshi. 1990 Dec;94(12):1103-21. [PubMed:1707214 ]
- Ouwerkerk-Mahadevan S, Mulder GJ: Inhibition of glutathione conjugation in the rat in vivo by analogues of glutathione conjugates. Chem Biol Interact. 1998 Apr 24;111-112:163-76. [PubMed:9679552 ]
- Mechanic GL: The qualitative and quantitative crosslink chemistry of collagen matrices. Adv Exp Med Biol. 1977;86B:699-708. [PubMed:20755 ]
- Okuno E, Tsujimoto M, Nakamura M, Kido R: 2-Aminoadipate-2-oxoglutarate aminotransferase isoenzymes in human liver: a plausible physiological role in lysine and tryptophan metabolism. Enzyme Protein. 1993;47(3):136-48. [PubMed:8087205 ]
- Takechi T, Okada T, Wakiguchi H, Morita H, Kurashige T, Sugahara K, Kodama H: Identification of N-acetyl-alpha-aminoadipic acid in the urine of a patient with alpha-aminoadipic and alpha-ketoadipic aciduria. J Inherit Metab Dis. 1993;16(1):119-26. [PubMed:8487492 ]
- Wu HQ, Ungerstedt U, Schwarcz R: L-alpha-aminoadipic acid as a regulator of kynurenic acid production in the hippocampus: a microdialysis study in freely moving rats. Eur J Pharmacol. 1995 Jul 25;281(1):55-61. [PubMed:8566117 ]
- Jung K, Reszka R, Kamlage B, Bethan B, Stephan C, Lein M, Kristiansen G: Tissue metabolite profiling identifies differentiating and prognostic biomarkers for prostate carcinoma. Int J Cancer. 2013 Dec 15;133(12):2914-24. doi: 10.1002/ijc.28303. Epub 2013 Jun 29. [PubMed:23737455 ]
- Danhauser K, Sauer SW, Haack TB, Wieland T, Staufner C, Graf E, Zschocke J, Strom TM, Traub T, Okun JG, Meitinger T, Hoffmann GF, Prokisch H, Kolker S: DHTKD1 mutations cause 2-aminoadipic and 2-oxoadipic aciduria. Am J Hum Genet. 2012 Dec 7;91(6):1082-7. doi: 10.1016/j.ajhg.2012.10.006. Epub 2012 Nov 8. [PubMed:23141293 ]
- Sell DR, Strauch CM, Shen W, Monnier VM: 2-aminoadipic acid is a marker of protein carbonyl oxidation in the aging human skin: effects of diabetes, renal failure and sepsis. Biochem J. 2007 Jun 1;404(2):269-77. [PubMed:17313367 ]
- Wijekoon EP, Skinner C, Brosnan ME, Brosnan JT: Amino acid metabolism in the Zucker diabetic fatty rat: effects of insulin resistance and of type 2 diabetes. Can J Physiol Pharmacol. 2004 Jul;82(7):506-14. [PubMed:15389298 ]
- Yuan W, Zhang J, Li S, Edwards JL: Amine metabolomics of hyperglycemic endothelial cells using capillary LC-MS with isobaric tagging. J Proteome Res. 2011 Nov 4;10(11):5242-50. doi: 10.1021/pr200815c. Epub 2011 Oct 17. [PubMed:21961526 ]
- Zeitoun-Ghandour S, Leszczyszyn OI, Blindauer CA, Geier FM, Bundy JG, Sturzenbaum SR: C. elegans metallothioneins: response to and defence against ROS toxicity. Mol Biosyst. 2011 Aug;7(8):2397-406. doi: 10.1039/c1mb05114h. Epub 2011 Jun 7. [PubMed:21647514 ]
- Sell DR, Strauch CM, Shen W, Monnier VM: Aging, diabetes, and renal failure catalyze the oxidation of lysyl residues to 2-aminoadipic acid in human skin collagen: evidence for metal-catalyzed oxidation mediated by alpha-dicarbonyls. Ann N Y Acad Sci. 2008 Apr;1126:205-9. doi: 10.1196/annals.1433.065. [PubMed:18448817 ]
- Wang TJ, Ngo D, Psychogios N, Dejam A, Larson MG, Vasan RS, Ghorbani A, O'Sullivan J, Cheng S, Rhee EP, Sinha S, McCabe E, Fox CS, O'Donnell CJ, Ho JE, Florez JC, Magnusson M, Pierce KA, Souza AL, Yu Y, Carter C, Light PE, Melander O, Clish CB, Gerszten RE: 2-Aminoadipic acid is a biomarker for diabetes risk. J Clin Invest. 2013 Oct;123(10):4309-17. doi: 10.1172/JCI64801. Epub 2013 Sep 16. [PubMed:24091325 ]
- Lin H, Levison BS, Buffa JA, Huang Y, Fu X, Wang Z, Gogonea V, DiDonato JA, Hazen SL: Myeloperoxidase-mediated protein lysine oxidation generates 2-aminoadipic acid and lysine nitrile in vivo. Free Radic Biol Med. 2017 Mar;104:20-31. doi: 10.1016/j.freeradbiomed.2017.01.006. Epub 2017 Jan 6. [PubMed:28069522 ]
- Elshenawy S, Pinney SE, Stuart T, Doulias PT, Zura G, Parry S, Elovitz MA, Bennett MJ, Bansal A, Strauss JF 3rd, Ischiropoulos H, Simmons RA: The Metabolomic Signature of the Placenta in Spontaneous Preterm Birth. Int J Mol Sci. 2020 Feb 4;21(3). pii: ijms21031043. doi: 10.3390/ijms21031043. [PubMed:32033212 ]
|
|---|